Background on the Sites and Samples
We first learned about the demographics of the first farmers across the Levant and Zagros from Lazaridis et al. (2016). From that work, we saw that there were at least two independent developments towards agriculture, with two distinct populations (Lazaridis et al., 2016). One was located in the Levant (Natufians), while the other was at Ganj Dareh, in Iran (Lazaridis et al., 2016). We learned from the paper that Natufians and Ganj Dareh were both nearly equal in Basal Eurasian, yet Ganj Dareh had a lot of ancestry from a lineage related to ANE, and Natufians had little to none of this ancestry (Lazaridis et al., 2016). The Levant Neolithic samples from PPNB to PPNC were a mix of something related to Natufians, and another lineage related to Anatolian farmers from Barcin and Mentese (Mathieson et al., 2015; Lazaridis et al., 2016).
Also in 2016, we caught our first glimpse of the Early Neolithic in Central Anatolia (Kilinc et al., 2016), on the Konya Plain. The Boncuklu group was actually a very early Neolithic group that did have small-scale agriculture (Baird et al., 2012), but was not “aceramic” as many believe. There were very early styles of pottery production and limited use of ceramics at this site; some of the earliest in West Asia (Fletcher et al., 2017). The Boncuklu farmers were a group with long runs of homozygosity, comparable to Western Hunter Gatherers (WHG) (Kilinc et al., 2016). Despite this, the Boncuklu samples are actually quite similar to the later farmers at Barcin, Mentese, and early European farmers (Kilinc et al., 2016), pointing to Anatolia as a potentially third place of independent farming production, along with the Levant and Zagros.
Broushaki et al. (2016) also brought us more samples from the Zagros, that clustered with those samples from Ganj Dareh. These samples were from Tepe Abdul Hosein and Wezmeh Cave (Broushaki et al., 2016).
When it came to actually looking at the ancestral breakdown of Anatolians, Lazaridis et al. (2016) came up with a very solid model where Anatolians were a mix of lineages related to Ganj Dareh, Levant Neolithic, and WHG, with mixture proportions of 0.387, 0.339, and 0.274, respectively.
For the purpose of this post, I am going to focus on Boncuklu and their role in the formation of the Anatolian Neolithic. Since Boncuklu capture method involved shotgun data, I am also using shotgun sequenced genomes from Broushaki et al. (2016) to use as my Iranian farmers (Tepe Abdul Hosein), and the Barcin farmers (Bar8, Bar31). Also, Kostenki14 and MA1 are being used as hunters, as these two form two poles of variation, and are important in the formation of WHG and EHG. Lineages related to both are also important to the formation of these farmers.
In the analysis of the Boncuklu and Barcin farmers, Kilinc et al. (2016) stated that diversity from mixing with other similar groups and/or admixture from more southern groups is possible in the transition from Boncuklu to the later Neolithic of Anatolia. Taking a look at various methods of analysis, it does seem quite possible that a more southern source did admix into the Anatolian farmers of Barcin Hoyuk.
D-stats
First, looking at D-stats, there isn’t anything that is very significant, in determining if Barcin or Boncuklu share more with any particular population. There are near significant results for Natufians and less for the Levant Neolithic samples.
| Outgroup | Test | Pop1 | Pop2 | f4 | Z-score | SNPs |
| Mbuti_DG | Ust_Ishim | Boncuklu | Anatolia_N1 | -0.000282 | -0.628 | 1081252 |
| Chimp | Ust_Ishim | Boncuklu | Anatolia_N1 | -0.000416 | -0.876 | 1036168 |
| Mbuti_DG | Natufian | Boncuklu | Anatolia_N1 | 0.00148 | 2.878 | 494327 |
| Chimp | Natufian | Boncuklu | Anatolia_N1 | 0.001535 | 2.843 | 472808 |
| Mbuti_DG | Levant_N | Boncuklu | Anatolia_N1 | 0.000891 | 2.171 | 804093 |
| Chimp | Levant_N | Boncuklu | Anatolia_N1 | 0.000694 | 1.556 | 769663 |
| Mbuti_DG | Tepe_Abdul | Boncuklu | Anatolia_N1 | -0.000556 | -1.448 | 1003898 |
| Chimp | Tepe_Abdul | Boncuklu | Anatolia_N1 | -0.000645 | -1.559 | 962310 |
| Mbuti_DG | Wezmeh | Boncuklu | Anatolia_N1 | -0.00025 | -0.562 | 1082448 |
| Chimp | Wezmeh | Boncuklu | Anatolia_N1 | -0.000415 | -0.882 | 1037321 |
| Mbuti_DG | CHG | Boncuklu | Anatolia_N1 | -0.000361 | -0.867 | 1083101 |
| Chimp | CHG | Boncuklu | Anatolia_N1 | -0.000445 | -1.031 | 1037961 |
| Mbuti_DG | Iron_Gates | Boncuklu | Anatolia_N1 | -0.000623 | -1.69 | 1076193 |
| Chimp | Iron_Gates | Boncuklu | Anatolia_N1 | -0.000712 | -1.803 | 1031346 |
| Mbuti_DG | EHG | Boncuklu | Anatolia_N1 | -0.000393 | -0.989 | 995924 |
| Chimp | EHG | Boncuklu | Anatolia_N1 | -0.000544 | -1.281 | 954789 |
| Mbuti_DG | W_Siberia | Boncuklu | Anatolia_N1 | 0.000176 | 0.404 | 818559 |
| Chimp | W_Siberia | Boncuklu | Anatolia_N1 | 0.000181 | 0.397 | 783377 |
| Mbuti_DG | Ami | Boncuklu | Anatolia_N1 | -0.001048 | -1.936 | 734487 |
| Chimp | Ami | Boncuklu | Anatolia_N1 | -0.001122 | -1.945 | 702374 |
f3-ratio
Also, looking at f3, there isn’t a lot to really support much in the way of admixture from Levant or Natufians into the Barcin Hoyuk samples.
| Pop1 | Pop2 | Target | f3 | std error | Z-score | SNPs |
| Boncuklu | Tepe_Abdul | Anatolia_N1 | 0.004 | 0.002013 | 1.987 | 494742 |
| Boncuklu | Wezmeh | Anatolia_N1 | 0.002506 | 0.002218 | 1.13 | 486870 |
| Boncuklu | Levant_N | Anatolia_N1 | -0.000089 | 0.001916 | -0.047 | 382114 |
| Boncuklu | Tepecik | Anatolia_N1 | 0.004217 | 0.00194 | 2.174 | 361064 |
| Boncuklu | Natufian | Anatolia_N1 | -0.001456 | 0.00235 | -0.619 | 222845 |
qpAdm Models
Switching over to qpAdm, the first thing to check is what Boncuklu is best modeled as. For the right populations the simplest and seemingly best for using Kostenki, Natufians, and Tepe_Abdul_Hosein_N as the left pops turned out to be Mbuti_DG, Ust_Ishim, GoyetQ116-1, Vestonice16, MA1, Ami, Onge, and Karitiana.
| probability | Natufian | Tepe_Abdul | Kostenki14 | std error | std error | std error |
| 0.236 | 0.232 | 0.449 | 0.32 | 0.142 | 0.107 | 0.074 |
While the probability and standard errors are a little high, it is hard to keep them any lower. While adding something like Ganj_Dareh_N or Levant_N will reduce the standard error, the probability also dips significantly.
Moving onto Anatolia_N1 (Bar31, Bar8), the right pops, or outgroups became Mbuti_DG, Ust_Ishim, Kostenki14, MA1, Natufian, Tepe_Abdul_Hosein_N, Ami, Onge, Karitiana.
| probability | Boncuklu | Levant_N | std error | std error |
| 0.815 | 0.687 | 0.313 | 0.064 | 0.064 |
Also, swapping Natufian and Levant_N between left and right populations:
| probability | Boncuklu | Natufian | std error | std error |
| 0.302 | 0.771 | 0.229 | 0.071 | 0.071 |
qpGraph Models
After seeing some decent fits in qpAdm, I tried to use qpGraph and see how easily they could be replicated. To being with, I started with a simple tree that contained just the Natufians and Tepe_Abdul_Hosein_N. The Interestingly, CHG comes out as a mix of Natufian, Tepe_Abdul_Hosein, and MA1. first tree should what would be expected, with Natufians and Tepe_Abdul_Hosein_N having similar amounts of Basal Eurasian, and with Iranian farmers having ancestry from a lineage related to MA1.
6/4/18 New Update: Simple qpGraphs
The purpose of these simple graphs is to avoid a lot of admixture edges and to show the steps in finding the ancestry of Boncuklu, Levant, and Barcin samples. Samples that are not absolutely needed, such as Ust_Ishim, MA1, CHG, and any ENA population will be left out. This also shows the work process in how I find where to look at for admixture events.
The first graph here shows that by branching Boncuklu off of Natufian, there is a significant Z-score with Iranian farmers. The next step will be to branch from Iranians to Boncuklu.

The next output had a worst Z-score that involved Kostenki, meaning that an admixture line from the Kostenki branch will be the next admixture source into Boncuklu.

After adding the admixture edge from near Kostenki, Boncuklu comes out mostly Kostenki-related, with the rest being a near even mix of Tepe Abdul Hosein and Natufian.
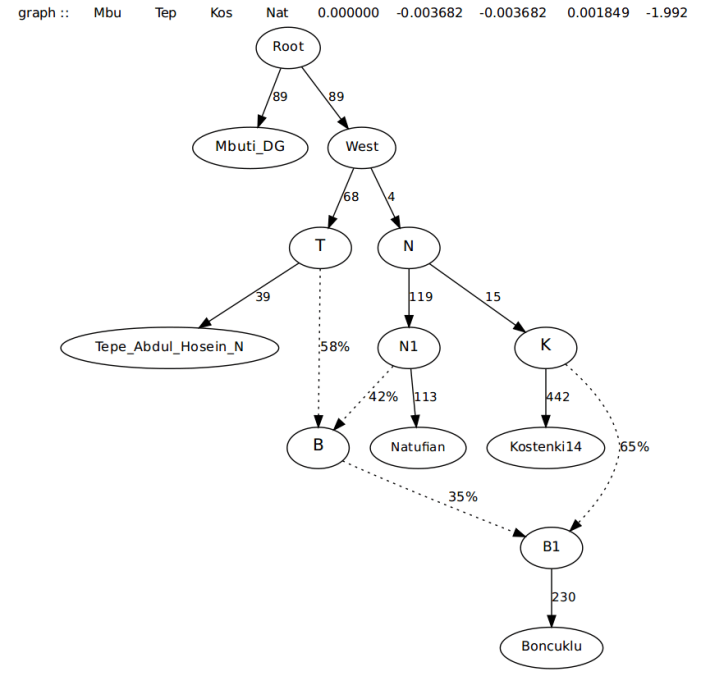
This created a pretty solid fit, so, the next step would be to see how the graph works with Adding Levant_N. Since the Levant samples harbor a lot of Natufian-related ancestry, that is where they will first be branched off of.

This output shows that an admixture edge is needed from a node on the Boncuklu line to Levant_N in order to drop the Z-score. A shared drift edge with Boncuklu was needed to move the worst Z-score closer to 2.

For the next run, I added the two shotgun samples from Barcin (Bar8, Bar31) to the tree.This left a worst Z > 3, so, the next step will be an edge from a node related to Natufians into Anatolia_N1.

For the last graph, the edge from a Natufian-related node admixes into the Barcin Hoyuk samples. There remains a worst Z > 2.5, that appears to be asking for Iranian admixture into the Levant_N population. While it isn’t necessary, I did look at adding an admixture edge from the Iranian farmers and it only added 2% admixture. This graph should be sufficient enough for a simple tree that incorporates all of the first farmers.
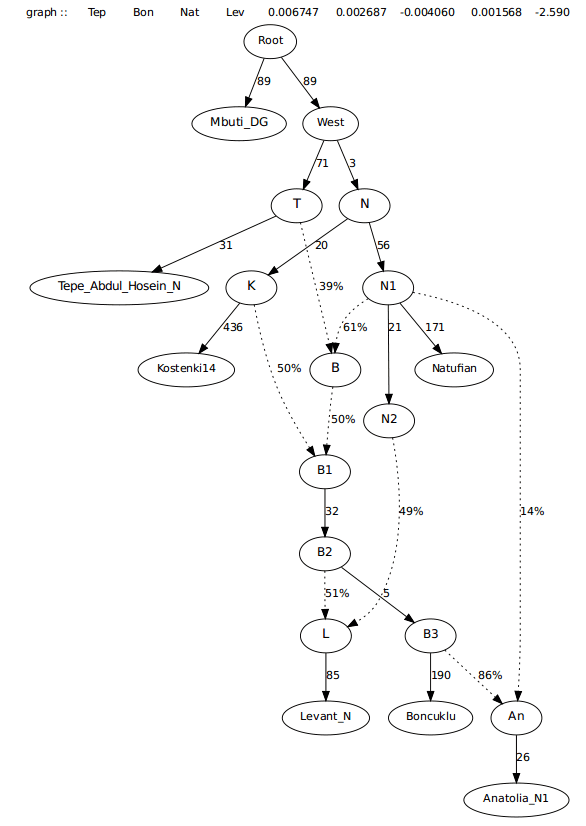
Below are more complex graphs, with an Eastern non-African source (Onge), Ust-Ishim and also MA-1.
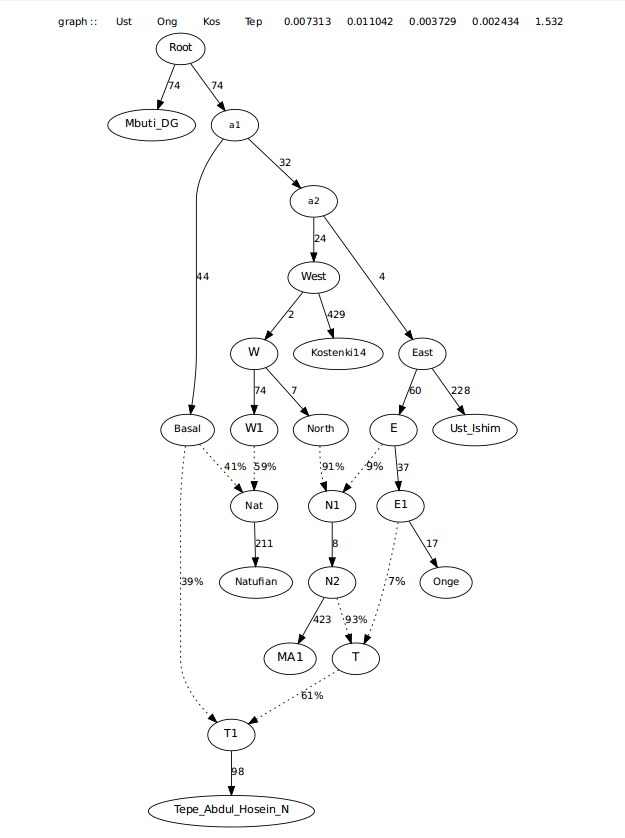
For the next step, I also added CHG to the tree, just to see where it was placed and to add another admixture source in West Asia. As other stats show, CHG was less basal than Iranian farmers, but also shifted towards Natufians, potentially sharing a formation story with Boncuklu.
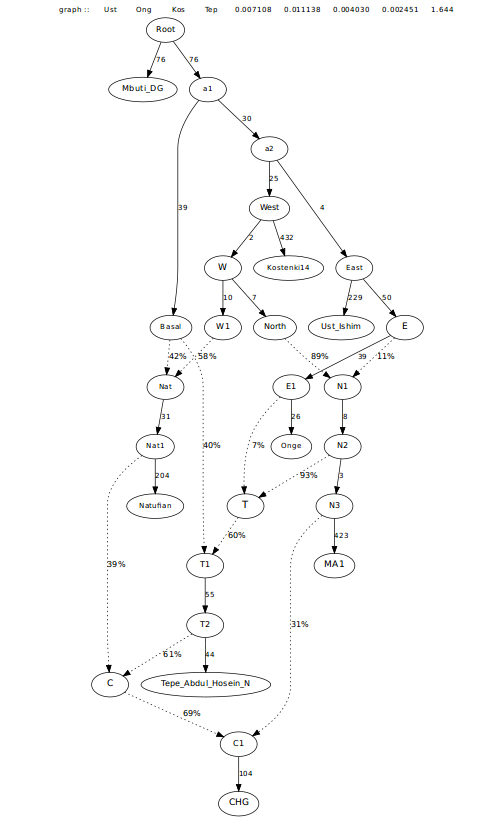
After achieving a good fit with CHG, I then went on to see just how Boncuklu fit into the picture. As expected, Boncuklu is best fit as a mix of Natufian, Iranian, and a lineage related to Kostenki14, with admixture also from CHG. Trying to do just Iranian or just CHG led to significantly higher Z-scores. Both were needed. It appears there is additional ANE in Boncuklu that is not accounted for in Iranians.

After applying a good fit for Boncuklu, I next wanted to see just how the farmers at Barcin Hoyuk fit into tree. For this, I first created a new simplified tree to use and also co-fit the Levant_N samples to see if the best fit includes additional southern ancestry into the later Northwest Anatolian Neolithic.

I also tried a run with just transversion sites, just to see how that would turn out.
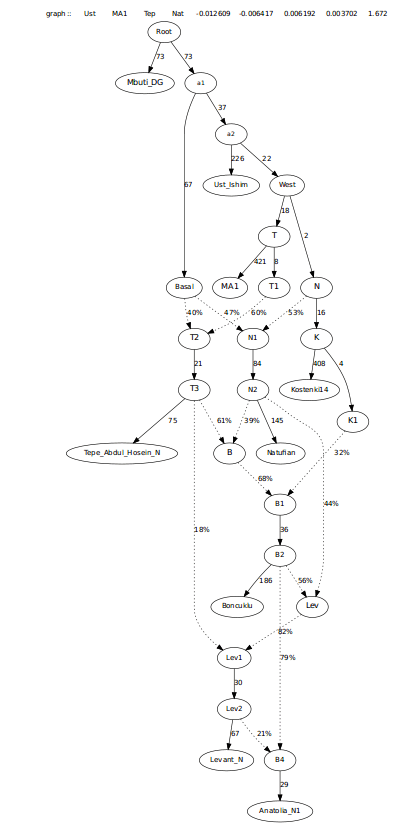
Just as a precaution, I ran another test, with Levant_N receiving admixture from a separate branch related to Boncuklu, that is ancestral to Anatolia_N1, to see if that reduced or eliminates the need for admixture from Levant_N. As it turns out, Anatolia then asked first for admixture from Natufians, rather than Levant_N. So, the question then becomes, was there another population that was related to Levant_N which lacked or had less Iranian farmer ancestry and mixed into the ancestors of the Barcin and European farmers? More samples, specifically from southern and especially southeast Anatolia, and PPNA will provide the answers which are needed.For argument sake, I tried both the Natufian and Levant_N admixture edges to see how they turned out. Having more a higher f4 and f3 with Natufians, rather than Levant, leads me to believe this is possible.


Conclusions
The end result here has the Barcin samples as a mix of Boncuklu and Levant_N (or a more Natufian-like group), in agreement with qpAdm. However, this output is only about two-thirds as much Levantine flow into later Anatolian farmers as qpAdm showed. Transversion sites placed more ancestry related to Iranian farmers into both Boncuklu and Levant_N. Boncuklu became significantly more Iranian than the previous run. The amount of flow from Levant_N into Barcin held pretty steady. While more samples across time and space, with similar capture methods, will help this to be resolved. There aren’t much in the way of D-stats or f3_ratio to suggest there is a lot of gene flow there. qpAdm and qpGraph do provide a strong case of gene-flow into Northwest Anatolia.
In upcoming posts I will look at the formation of European farmers, hunter admixture into the Middle Neolithic, and also at the roots of Epipaleolithic and Mesolithic European hunters.
References
Baird D. The Late Epipaleolithic, Neolithic, and Chalcolithic of the Anatolian Plateau, 13,000–4000 BC. In: Potts D., editor. A Companion to the Archaeology of the Ancient Near East. Wiley-Blackwell; 2012. pp. 431–466.
Broushaki F., Thomas M.G., …, Berger J. Early Neolithic genomes from the eastern Fertile Crescent. Science. 2016; 353: pp. 499-503.
Fletcher A., Baird D., Spataro M. & Fairbairn A. Early ceramics in Anatolia: Implications for the production and use of the earliest pottery. The evidence from Boncuklu Hoyuk. Cambridge Archaeological Journal. 2017; 27 (2): pp. 351-369.
Kilinc G.M., Omrak A., …, Gotherstrom A. The demographic development of the first farmers in Anatolia. Current Biology. 2016; 26(19): pp. 2659–2666.
Lazaridis I., Nadel D., …, Reich D. Genomic insights into the origin on farming in the ancient Near East. Nature. 2016;536: pp. 419-424.
Mathieson I., Lazaridis I., Rohland N., Mallick S., Patterson N., Roodenberg S.A., Harney E., Stewardson K., Fernandes D., Novak M. Genome-wide patterns of selection in 230 ancient Eurasians. Nature. 2015;528:499–503. [PubMed]
Go ahead and post any comments or questions!
LikeLike
Great to see you’ll have this blog to post all your work. I might not be commenting much (decided to have my own place too where my comments won’t bother anyone), but I’ll surely be following this and giving credit whenever I comment about anything posted here.
Looking forward to all the upcoming posts. Good luck and thanks for all the work!
LikeLike
Thanks, Alberto! Best of luck to you too.
LikeLike
Hi Alberto, whats your blog’s URL please?
LikeLike
adnaera.com
LikeLike
I like your analysis, will follow your future posts, I’m very interested in the deep history of Anatolian farmers and European hunters.
Looking forward to your analysis of the Villabruna cluster and it’s affinity to Ancient Near Eastern groups.
LikeLike
Thank you! I’ll have more posts coming soon.
LikeLike
A new post will be coming in the next couple of days. It will potentially be one regarding a split with Danubian and Mediterranean groups, and then a larger project focusing on the Danubian groups. Specifically, I plan on looking at the formation of the Linear Pottery Culture (LBK) and the many related groups. Do they have Vinca ancestry as some have suggested, or is it merely culturally influence (pots, not people)? We shall see!
LikeLike
Very thorough analysis Chad.
Seems to correlate nicely with archaeology.
It would be great to see some Epipaleolthic (pre 10000 BC) genomes from Anatolia, like the recently uncovered from Pinarbasi.
Any guestimates on its affinities ?
LikeLike
Due to the lithics, other cultural materials, and low genetic diversity, I think they are old. The 15-17kya samples might be surprisingly like Boncuklu.
LikeLike
I have cleaned up the post a little and hopefully made it a little more reader-friendly. Thanks to Rui Martiniano for the suggestions. I will update this with some more visually appealing stuff, such as maps too.
LikeLike
As you hint, I do think there should be some yet upsampled group that existed around southeastern Anatolia / northern Levant, probably isolated for quite a while but somehow became instrumental in early agriculture and blew up. A few disjointed speculative thoughts on this:
Lazardis 2016 stated that the HG portion of Anatolians (Barcin) was “more west” than WHG — let’s stick to calling them UHG. I recall it was also proposed that European WHGs somehow interacted with eastern non-Africans, though I’m unsure what the consensus is about this now… so UHGs not participating in this interaction would explain them being “more western.”
I’m curious whether G2a could be the Y-hg associated with these hypothetical UHG. It’s strange how it’s not seen anywhere until neolithic farming, yet, it’s also not found in Natufians and other Levantines. (Prior to the DNA data I would’ve guessed to see it there given it’s where there’s the most evidence for the earliest proto-agriculture). G2a generally deserves more attention… obviously it’s a big part of the story and its ultimate origins are still unknown.
Perhaps obsidian mining/trading plays a role here too. I can imagine a scenario where a Kostenki-related UHG obsidian people and a Basal-related farming people coalesce around southeastern Anatolia / northern Levant to form the proto-Anatolians. Another case of female exogamy at the border? 😉
LikeLike
Haplogroup G I believe has origins in the Zagros mountains, G1a, G2b, and G2a1 are all found in the Iranian Neolithic, Anatolian Neolithic has a major part of their ancestry from an Iran_Neo like population, as per this post analysis and Lazaridis et al(2016).
LikeLike
That is, of course, very possible too.
LikeLike
PF,
Which part of Lazaridis (2016) are you seeing that part of the HG in Barcin being less ANE or ENA than WHG? I’m not seeing it here.
LikeLike
It’s mentioned in Supplementary Info section 7:
“…we give an interpretation of the origin of the Anatolian Neolithic that interprets the negative EHG mixture proportion (Table S7.3) in terms of admixture (into the Anatolian Neolithic) of a population that is not exactly WHG, but rather residing on the EHG→WHG cline beyond WHG.”
And in section 10, where they try creating ghost populations:
“Notice that the proportions from the “Ghost” population are fairly stable at around ~1/3, and this population is inferred to be (based on the λ parameter) beyond both WHG and Switzerland_HG on the respective EHG→WHG and EHG→Switzerland_HG clines and closer to Switzerland_HG than to WHG. Future sampling may reveal whether such a population, whose existence is hypothesized here as a way to better model the Anatolian Neolithic, did actually exist.”
LikeLike
Yes, I see. I think that is possible. The more complex model doesn’t take a lot more ANE or ENA than that found in Ganj Dareh. There’s a need for some epipaleolithic samples to sort this out.
LikeLike